|
10/31/2023 0 Comments Rstudio install package
ĭrwxr-xr-x 2 root root 36K Apr 14 06:13 binĭrwxr-xr-x 6 root root 4.0K Nov 10 10:43 includeĭrwxr-xr-x 86 root root 4.0K Mar 18 08:24 libĭrwxr-xr-x 2 root root 4.0K Mar 2 06:54 lib64ĭrwxr-xr-x 3 root root 4.0K Mar 18 08:24 libexecĭrwxr-xr-x 2 root root 20K Apr 11 06:22 sbinĭrwxr-xr-x 122 root root 4.0K Mar 18 08:24 shareĭrwxr-xr-x 8 root root 4.0K Apr 2 06:06 srcĪs you can see they are all owned by root and have no write privileges for anybody else. For example, on one of my work boxes, here are all the directories under /usr/ $ lsl -la /usrĭrwxr-xr-x 14 root root 4.0K Mar 10 12:05. I have never previously encountered this issue with Bioconductor on other machines (Mac, Linux/PopOS 20.04) I have only successfully installed BiocManager and BiocVersion but no packages. I also tried re-installing BiocManager as suggested - no change, but this was a fresh install to begin with. I tried the ghost-binary-repo code but it is no longer posted there. those directories all exist in that location. So if I try getOption > getOption("repos") Installation paths not writeable, unable to update packagesĬluster, Matrix, mgcv, nlme, spatial, survival 'getOption("repos")' replaces Bioconductor standard repositories, seeīioconductor version 3.14 (BiocManager 1.30.16), R 4.1.3 () I get the following > BiocManager::install() 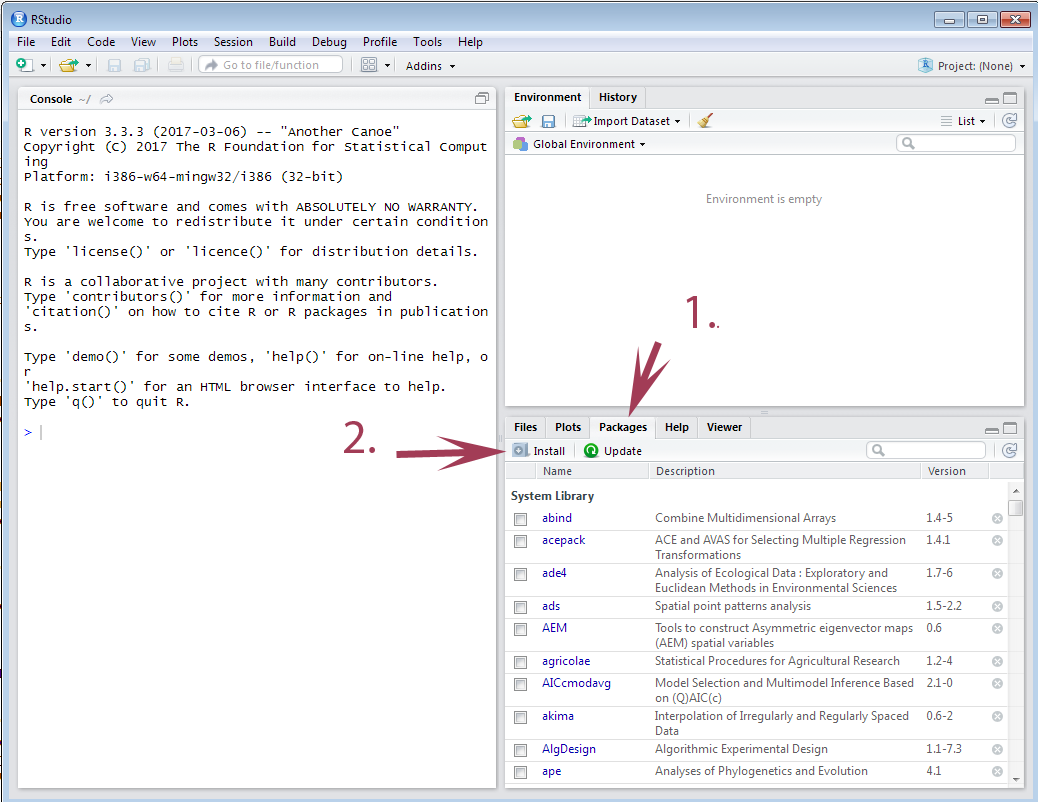
That is a fresh install on a new computer - no updating. I am also having this issue installing Bioconductor on fresh new Linux (x86_64-pc-linux-gnu (64-bit), PopOS 21.10) R is 4.1.3.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
